Fahlbusch 2022 Int J Mol Sci
| Fahlbusch P, Nikolic A, Hartwig S, Jacob S, Kettel U, Köllmer C, Al-Hasani H, Lehr S, Müller-Wieland D, Knebel B, Kotzka J (2022) Adaptation of oxidative phosphorylation machinery compensates for hepatic lipotoxicity in early stages of MAFLD. Int J Mol Sci 23:6873. https://doi.org/10.26124/10.3390/ijms23126873 |
Fahlbusch P, Nikolic A, Hartwig S, Jacob S, Kettel U, Koellmer C, Al-Hasani H, Lehr S, Mueller-Wieland D, Knebel B, Kotzka J (2022) Int J Mol Sci
Abstract: Alterations in mitochondrial function are an important control variable in the progression of metabolic dysfunction-associated fatty liver disease (MAFLD), while also noted by increased de novo lipogenesis (DNL) and hepatic insulin resistance. We hypothesized that the organization and function of a mitochondrial electron transport chain (ETC) in this pathologic condition is a consequence of shifted substrate availability. We addressed this question using a transgenic mouse model with increased hepatic insulin resistance and DNL due to constitutively active human SREBP-1c. The abundance of ETC complex subunits and components of key metabolic pathways are regulated in the liver of these animals. Further omics approaches combined with functional assays in isolated liver mitochondria and primary hepatocytes revealed that the SREBP-1c-forced fatty liver induced a substrate limitation for oxidative phosphorylation, inducing enhanced complex II activity. The observed increased expression of mitochondrial genes may have indicated a counteraction. In conclusion, a shift of available substrates directed toward activated DNL results in increased electron flows, mainly through complex II, to compensate for the increased energy demand of the cell. The reorganization of key compounds in energy metabolism observed in the SREBP-1c animal model might explain the initial increase in mitochondrial function observed in the early stages of human MAFLD.
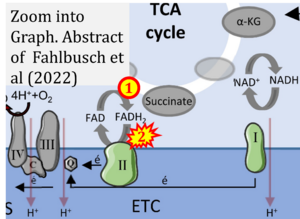
Correction: FADH2 and Complex II
- FADH2 is shown as the substrate feeding electrons into Complex II (CII). This is wrong and requires correction - for details see Gnaiger (2024).
- Gnaiger E (2024) Complex II ambiguities ― FADH2 in the electron transfer system. J Biol Chem 300:105470. https://doi.org/10.1016/j.jbc.2023.105470 - »Bioblast link«
Labels:
Enzyme: Complex II;succinate dehydrogenase