Template talk:Next oroboros ecosystem agenda: Difference between revisions
(→Next) |
Tindle Lisa (talk | contribs) |
||
| Line 22: | Line 22: | ||
== Bioblast alert 2021 == | == Bioblast alert 2021 == | ||
Prep for circular: | Prep for circular: | ||
== MitoPedia alert == | == MitoPedia alert == | ||
Revision as of 14:31, 16 September 2021
- Antunes Diana (talk) 08:58, 5 May 2020 (CEST) Format for new Agenda/Alerts entries:
{|
| logos
| title in bold <br>
- link
|}
e.g.,


Did you know that the proper use of TIP2k-Filter Papers may avoid an unstable O2 slope neg. signal during Instrumental background correction?
- »Instrumental background« and »TIP2k Filter Papers«
- See the scheme:
Bioblast alert 2021
Prep for circular:
MitoPedia alert
Next
O2k-Brief alert
Next
O2k-Publications alert
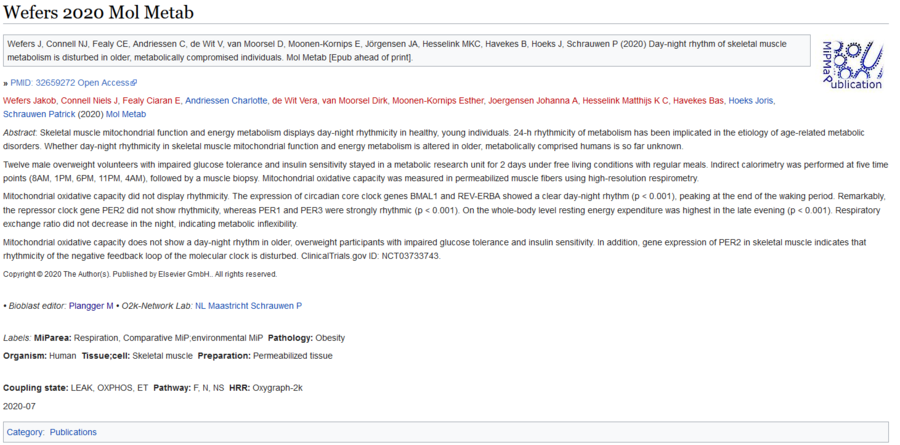
Author names of O2k-Publications/ Brief for the Agenda vs Bioblast
- 1.Oroboros Ecosystem Agenda: SOP: Surname with long version and First name with short version. Please, see: Template alerts.
- Example: Oroboros Ecosystem Agenda example
- This means that we have to edit manually the author names within the O2k-Publications/ Brief alerts (i.e., Antunes Diana (from Bioblast) to Antunes D (to Agenda entries)) if they are long or copy from the gray box.
- 2.Bioblast: SOP: [1] – examples are available using both Surname and First name with long version (e.g., Antunes Diana). If in Bioblast the authors are with short surname in the links please update it to complete surname, guidelines are below.
- 1.Oroboros Ecosystem Agenda: SOP: Surname with long version and First name with short version. Please, see: Template alerts.
Guidelines to update a reference
- When creating an O2k-Publication/Brief alert, it is the responsibility of the author to check if the reference is accordingly to the correct guidelines:
- Correct format for a person called “First Name Surname”:
- In the gray box should be “Surname FN” - From where references to be used as O2k-Publication/Brief alerts are copied from.
- In the links should be “ Surname First N”
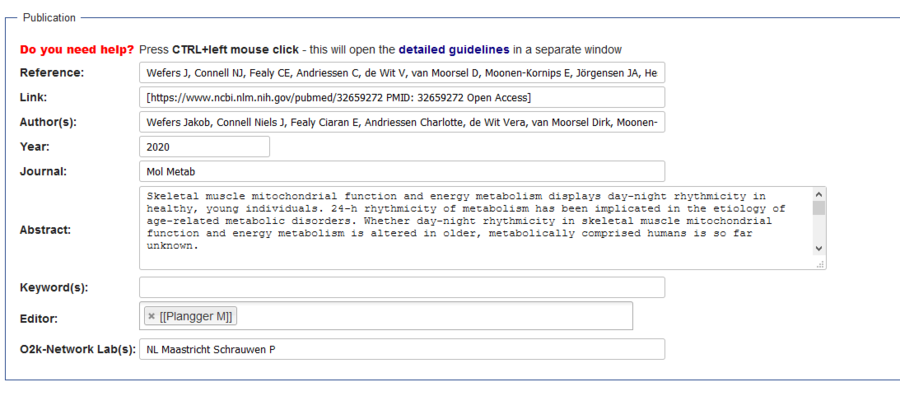
- How to edit:
- A) Check if all authors with profiles in Bioblast (blue links) have their profiles with full name. If they have, proceed to next step (B). If not, you need to move it following the steps “How to move a profile” below.
- B) Click in “Edit with form”.
- C) The gray box is edited in “Reference” and the links are changed in “Author(s)”. Change it accordingly and click in “save page” at the end of the page.
- How to edit:
How to move a profile (instructions sent by Marija):
So let’s take for example my page: Beno M [2]
- As you can see there is a redirect from Beno Marija to Beno M. What has to be done is to switch the content of those pages. Here are the steps:
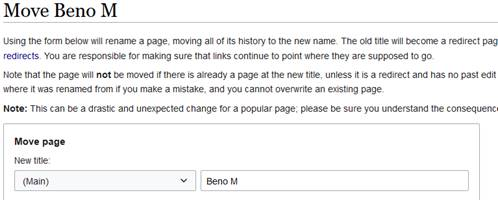
- 1. Go to e.g. “Beno M” on edit
- 2. Click on “Move”
- 3. Change the new title to the full name of the person. In my case it will be Beno M to Beno Marija
- As you can see there is a redirect from Beno Marija to Beno M. What has to be done is to switch the content of those pages. Here are the steps:
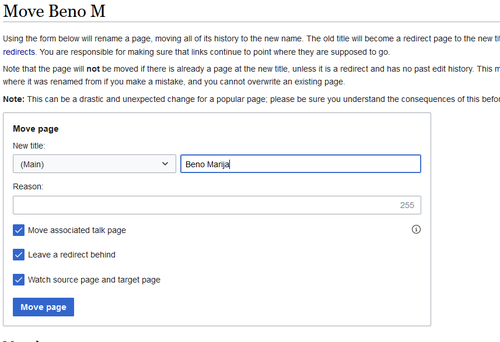
- 4. Now it should look like that:
- 5. Click on “Move” and leave all the ticked boxes
- 6.Done
- After that you can continue in step B.
- If you have any doubt you can contact Cristiane, Marija or Mario.
Next
O2k-Open Support alert
- While in Home Office: Gnaigere (talk) 14:03, 19 March 2020 (CET) https://wiki.oroboros.at/index.php/Talk:O2k-Open_Support_alert - Many labs will move to home office mode. In line with our special DL7.4 home office offer (see Agenda today), therefore, I suggest to post in our Agenda analysis-linked O2k-Open Support cases during this time, and collect real-time operation cases for future releases.
Next
- Sept-22


Check out the new excel templates for mitochondrial membrane potential analysis using fluorescence dyes
- » mt-membrane potential Excel templates« and »'MiPNet24.09 General Template for Mt-membrane Potential Analysis«
- communicated by »Komlodi Timea« and »Cardoso Luiza«
- Sept-29


Check out the new excel templates for H2O2 flux analysis with the Amplex UltraRed assay
- » H2O2 flux Excel templates« and »'MiPNet24.10 H2O2 flux analysis«
- communicated by »Komlodi Timea« and »Cardoso Luiza«
- Oct-06


Check out the new excel templates for ATP production analysis with the Magnesium Green
- »MgG Excel templates« and »MiPNet26.10 MgG data analysis«
- communicated by »Cardoso Luiza« and »Komlodi Timea«
This one needs a BioBlast link:
- Month-##


When filling in the volume of uncoupler in the DatLab mark or the excel template, should one enter the total volume added or the volume to get to the highest flux?
The volume added until the mark, equal to the highest flux, should be entered. Anything titrated after this mark should be summed to the other volumes before the next mark because this will be considered to calculate the sample dilution.
- »LINK and O2 flux analysis with DatLab 7.4«
- communicated by »Cardoso Luiza« and »[[ ]]«
Under discussion
Removed for correction
- Nov-30 Mo


Noisy oxygen signal? Check the OroboPOS and OroboPOS-Connector to clean electrical connections.
Contaminating grease, salt crystals and moisture can be removed with a dry paper cloth or a paper cloth moistened with water and then absolute ethanol.
- »Cleaning the electrical connections«
- communicated by »Schmitt Sabine« and »Komlodi Timea«
WGT recommended not to use water for cleaning. Erich introduced water for cleaning to remove salt crystals.


Do you perform the leaky chamber test after chamber assembly?
An unnoticed leak from the chamber may affect your data, therefore we recommend performing the Leaky chamber test - especially after chamber reassembly.
- »Leaky chamber test« and » https://intern.oroboros.at/index.php/Overnight_test_after_O2k-chamber_assembly«
O2k-Open Support alert circulars
In preparation
MitoFit Preprints
Promoting transparency in scientific contributions
MitoFit Preprints encourages Open Access to relevant scientific contributions of acknowledged persons
»Open Access Acknowledgements«
Data analysis templates and MipNets
For discussion
Here we discuss suggestions for the Oroboros Ecosystem Agenda 2020 (if the specific alert section is yet to be attributed)

Controlled stability of the MitoKit-CII based on HPLC and LCMS analysis: No degradation of MitoKit-CII compounds MitoKit-CII/Succinate-nv (NV118) and MitoKit-CII/Malonate-nv (NV161) for following storage conditions: - Proven -80 °C solid stability for more than 3 years (3 years 7 months)
- Proven -20 °C solid stability for 1 year
- Proven DMSO stability at room temperature for 1 week
- Proven DMSO stability at -20 °C for 6 months (ongoing)
2021
- 2021-01-06 https://www.jstor.org/stable/1706346?seq=1
Postponed events