- high-resolution terminology - matching measurements at high-resolution
Glutamate-anaplerotic pathway control state
Description
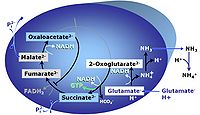
G: Glutamate is an anaplerotic NADH-linked type 4 substrate (N). When supplied as the sole fuel substrate in the glutamate-anaplerotic pathway control state, G is transported by the electroneutral glutamate-/OH- exchanger, and is oxidised via mt-glutamate dehydrogenase in the mitochondrial matrix. The G-pathway plays an important role in glutaminolysis.
Abbreviation: G
Reference: Gnaiger 2020 BEC MitoPathways
Communicated by Gnaiger E 2016-01-25, edited 2016-11-29.
G(L)
LEAK state (L) with G alone as substrate can be evaluated in the following SUIT protocol:
-
- DL-Protocol for permeabilized cells (pce): SUIT-007 O2 ce-pce D030
-
G(P)
OXPHOS state (P) with G alone as substrate can be evaluated in the following SUIT protocol:
-
- DL-Protocol for permeabilized cells (pce): SUIT-007 O2 ce-pce D030
-
G(E)
ET state (E) with G alone as substrate can be evaluated in the following SUIT protocol:
-
- DL-Protocol for permeabilized cells (pce): SUIT-007 O2 ce-pce D030
-
Details
- In human skeletal muscle mitochondria, OXPHOS capacity with glutamate alone (GP) is 50% to 85% of respiration in the GM-pathway control state (GMP). Accumulation of fumarate inhibits succinate dehydrogenase and glutamate dehydrogenase.
- Glutaminolysis: Glutamate derived from hydrolyzation of glutamine is an important aerobic substrate in cultured cells, supporting anaplerosis. Mitochondrial glutamate dehydrogenase is particularly active in astrocytes, preventing glutamate-induced neurotoxicity.
- Anaplerosis from malate: mtNAD-malic enzyme supports an anaplerotic pathway when carbohydrate is limiting.
SUITbrowser question: Glutamine/glutamate
- The SUITbrowser can be used to find the best SUIT protocols to analyse glutamine/glutamate metabolism, among other research questions.
MitoPedia concepts:
SUIT state